ASTM E3354-22
(Guide)Standard Guide for Application of Molecular Biological Tools to Assess Biological Processes at Contaminated Sites
Standard Guide for Application of Molecular Biological Tools to Assess Biological Processes at Contaminated Sites
SIGNIFICANCE AND USE
4.1 Contaminated sites subject to remediation are growing in complexity and associated remediation costs, presenting a challenge for managers of contaminated sites. The need to properly monitor, evaluate, and report remediation processes (including physical, chemical, and biological) characterizing site conditions and contaminant mass and attenuation is critical for the evaluation and selection of effective remediation strategies. Assessment and characterization of biological processes associated with contaminant attenuation is supported and improved by the accurate and consistent use of molecular biological tools (MBTs) including data acquisition, interpretation, and reporting.
4.2 The development of this guide through ASTM International is designed to meet the needs of managers of contaminated sites within the United States and elsewhere. The variety of available MBTs and the complexity with which they are currently being applied are not addressed in existing ASTM International Standards. The principal users of this guide should be industry project managers, regulators, consultants, analytical laboratories, and community stakeholders.
SCOPE
1.1 This guide provides a framework for the application of molecular biological tools (MBTs) to assess and characterize in-situ biological processes to improve contaminated soil and groundwater management. While the focus of this guide is on in-situ biological processes, some concepts of how to apply MBTs can also be applied to ex-situ bioremediation approaches (for example, biopiles, bioreactors) to support design, operation, and troubleshooting. The intent of this guide is to develop a consistent way in which MBTs are applied at contaminated sites, not to develop expertise. Technical experts need to be engaged whenscoping, planning, executing, and interpreting data for MBTs. Lastly, there is a brief description of isotopic techniques within section 5.2; however, the scope and focus of this guide is the use of nucleic acid-based MBTs to assess biological processes at contaminated sites.
1.2 This standard does not purport to address all of the safety concerns, if any, associated with its use. It is the responsibility of the user of this standard to establish appropriate safety, health, and environmental practices and determine the applicability of regulatory limitations prior to use.
1.3 This international standard was developed in accordance with internationally recognized principles on standardization established in the Decision on Principles for the Development of International Standards, Guides and Recommendations issued by the World Trade Organization Technical Barriers to Trade (TBT) Committee.
General Information
- Status
- Published
- Publication Date
- 30-Jun-2022
- Technical Committee
- E50 - Environmental Assessment, Risk Management and Corrective Action
- Drafting Committee
- E50.04 - Corrective Action
Relations
- Effective Date
- 10-Feb-2002
Overview
ASTM E3354-22: Standard Guide for Application of Molecular Biological Tools to Assess Biological Processes at Contaminated Sites provides environmental professionals a systematic framework for using molecular biological tools (MBTs) to assess in-situ biological processes in contaminated soil and groundwater. The growing complexity of contaminated sites and the need for effective, science-based remediation strategies have driven the adoption of MBTs. These tools, which include nucleic acid-based analyses such as PCR, qPCR, and next generation sequencing (NGS), enable better characterization of site conditions, support remediation decisions, and enhance the consistency of site assessments.
ASTM E3354-22 is intended for project managers, environmental consultants, regulators, analytical laboratories, and community stakeholders who are responsible for the management, monitoring, and reporting of remedial processes. By providing a consistent approach to MBT application at contaminated sites, this standard helps bridge technical gaps and promotes informed decision-making throughout the site lifecycle.
Key Topics
- Definition and Scope of MBTs: Outlines the use of nucleic acid-based molecular biological tools-and briefly, isotopic techniques-for identifying and characterizing microorganisms and key biological processes supporting contaminant attenuation.
- Framework for Application: Describes a stepwise approach to integrating MBTs in site assessment, remedy selection, implementation, and monitoring.
- Types of Molecular Tools:
- Polymerase Chain Reaction (PCR) and Quantitative PCR (qPCR): Detect presence and measure abundance of target functional genes or microorganisms associated with biodegradation.
- Next Generation Sequencing (NGS): Provides comprehensive profiling of microbial communities and their genetic capabilities.
- Omics Approaches: Includes genomics, transcriptomics, metagenomics, proteomics, and metabolomics for in-depth understanding of microbial function and activity.
- Isotopic Techniques (CSIA and SIP): Offer supporting evidence of contaminant transformation via stable isotope probing and analysis.
- Complementary Data Integration: Stresses the importance of linking MBT data with traditional hydrogeological, geochemical, and contaminant concentration data to develop conceptual site models (CSMs).
Applications
The practical applications of ASTM E3354-22 are broad and provide significant value across the contaminated site management lifecycle:
- Site Assessment: MBTs help identify key microorganisms or functional genes that drive contaminant biodegradation, allowing practitioners to determine if baseline conditions support natural attenuation processes.
- Remedy Design and Optimization: Data from MBTs guide remedial strategy selection, such as biostimulation or bioaugmentation, and help troubleshoot or refine ongoing remediation by pinpointing biological limiting factors.
- Performance Monitoring: Ongoing application of MBTs allows for direct monitoring of remediation effectiveness, serving as an early indicator of microbial activity prior to observable contaminant reductions.
- Stakeholder Communication: The use of standardized MBT approaches enhances transparency and supports regulatory compliance, strengthening stakeholder and community trust.
- Ex-situ Bioremediation Support: While focused on in-situ processes, the framework is also applicable to ex-situ methods (e.g., biopiles, bioreactors) to guide design, operation, and troubleshooting.
Related Standards
The application of ASTM E3354-22 is frequently supported by other standards and guidance documents, including:
- ASTM D6771: Practice for Low-Flow Purging and Sampling Used for Groundwater Monitoring.
- U.S. EPA Guidance Documents: Including protocols for sampling, laboratory analysis, and data considerations in microbial and isotope studies.
- EPA 2008 Guide: Assessing Biodegradation and Source Identification of Organic Ground Water Contaminants Using Compound Specific Isotope Analysis (CSIA).
- ASTM Committee E50: Environmental Assessment, Risk Management, and Corrective Action standards.
Conclusion
Implementing ASTM E3354-22 adds clarity, rigor, and consistency to the use of molecular biological tools in contaminated site assessment and remediation. The guide enhances decision-making by providing actionable data that support effective, targeted remediation strategies and regulatory confidence. For organizations seeking to optimize remediation investments while managing risks and communicating effectively with stakeholders, this standard represents an essential resource in the evolving field of environmental molecular diagnostics.
Buy Documents
ASTM E3354-22 - Standard Guide for Application of Molecular Biological Tools to Assess Biological Processes at Contaminated Sites
Frequently Asked Questions
ASTM E3354-22 is a guide published by ASTM International. Its full title is "Standard Guide for Application of Molecular Biological Tools to Assess Biological Processes at Contaminated Sites". This standard covers: SIGNIFICANCE AND USE 4.1 Contaminated sites subject to remediation are growing in complexity and associated remediation costs, presenting a challenge for managers of contaminated sites. The need to properly monitor, evaluate, and report remediation processes (including physical, chemical, and biological) characterizing site conditions and contaminant mass and attenuation is critical for the evaluation and selection of effective remediation strategies. Assessment and characterization of biological processes associated with contaminant attenuation is supported and improved by the accurate and consistent use of molecular biological tools (MBTs) including data acquisition, interpretation, and reporting. 4.2 The development of this guide through ASTM International is designed to meet the needs of managers of contaminated sites within the United States and elsewhere. The variety of available MBTs and the complexity with which they are currently being applied are not addressed in existing ASTM International Standards. The principal users of this guide should be industry project managers, regulators, consultants, analytical laboratories, and community stakeholders. SCOPE 1.1 This guide provides a framework for the application of molecular biological tools (MBTs) to assess and characterize in-situ biological processes to improve contaminated soil and groundwater management. While the focus of this guide is on in-situ biological processes, some concepts of how to apply MBTs can also be applied to ex-situ bioremediation approaches (for example, biopiles, bioreactors) to support design, operation, and troubleshooting. The intent of this guide is to develop a consistent way in which MBTs are applied at contaminated sites, not to develop expertise. Technical experts need to be engaged whenscoping, planning, executing, and interpreting data for MBTs. Lastly, there is a brief description of isotopic techniques within section 5.2; however, the scope and focus of this guide is the use of nucleic acid-based MBTs to assess biological processes at contaminated sites. 1.2 This standard does not purport to address all of the safety concerns, if any, associated with its use. It is the responsibility of the user of this standard to establish appropriate safety, health, and environmental practices and determine the applicability of regulatory limitations prior to use. 1.3 This international standard was developed in accordance with internationally recognized principles on standardization established in the Decision on Principles for the Development of International Standards, Guides and Recommendations issued by the World Trade Organization Technical Barriers to Trade (TBT) Committee.
SIGNIFICANCE AND USE 4.1 Contaminated sites subject to remediation are growing in complexity and associated remediation costs, presenting a challenge for managers of contaminated sites. The need to properly monitor, evaluate, and report remediation processes (including physical, chemical, and biological) characterizing site conditions and contaminant mass and attenuation is critical for the evaluation and selection of effective remediation strategies. Assessment and characterization of biological processes associated with contaminant attenuation is supported and improved by the accurate and consistent use of molecular biological tools (MBTs) including data acquisition, interpretation, and reporting. 4.2 The development of this guide through ASTM International is designed to meet the needs of managers of contaminated sites within the United States and elsewhere. The variety of available MBTs and the complexity with which they are currently being applied are not addressed in existing ASTM International Standards. The principal users of this guide should be industry project managers, regulators, consultants, analytical laboratories, and community stakeholders. SCOPE 1.1 This guide provides a framework for the application of molecular biological tools (MBTs) to assess and characterize in-situ biological processes to improve contaminated soil and groundwater management. While the focus of this guide is on in-situ biological processes, some concepts of how to apply MBTs can also be applied to ex-situ bioremediation approaches (for example, biopiles, bioreactors) to support design, operation, and troubleshooting. The intent of this guide is to develop a consistent way in which MBTs are applied at contaminated sites, not to develop expertise. Technical experts need to be engaged whenscoping, planning, executing, and interpreting data for MBTs. Lastly, there is a brief description of isotopic techniques within section 5.2; however, the scope and focus of this guide is the use of nucleic acid-based MBTs to assess biological processes at contaminated sites. 1.2 This standard does not purport to address all of the safety concerns, if any, associated with its use. It is the responsibility of the user of this standard to establish appropriate safety, health, and environmental practices and determine the applicability of regulatory limitations prior to use. 1.3 This international standard was developed in accordance with internationally recognized principles on standardization established in the Decision on Principles for the Development of International Standards, Guides and Recommendations issued by the World Trade Organization Technical Barriers to Trade (TBT) Committee.
ASTM E3354-22 is classified under the following ICS (International Classification for Standards) categories: 07.080 - Biology. Botany. Zoology. The ICS classification helps identify the subject area and facilitates finding related standards.
ASTM E3354-22 has the following relationships with other standards: It is inter standard links to ASTM D6771-02. Understanding these relationships helps ensure you are using the most current and applicable version of the standard.
ASTM E3354-22 is available in PDF format for immediate download after purchase. The document can be added to your cart and obtained through the secure checkout process. Digital delivery ensures instant access to the complete standard document.
Standards Content (Sample)
This international standard was developed in accordance with internationally recognized principles on standardization established in the Decision on Principles for the
Development of International Standards, Guides and Recommendations issued by the World Trade Organization Technical Barriers to Trade (TBT) Committee.
Designation: E3354 − 22
Standard Guide for
Application of Molecular Biological Tools to Assess
Biological Processes at Contaminated Sites
This standard is issued under the fixed designation E3354; the number immediately following the designation indicates the year of
original adoption or, in the case of revision, the year of last revision. A number in parentheses indicates the year of last reapproval. A
superscript epsilon (´) indicates an editorial change since the last revision or reapproval.
1. Scope 2.2 U.S. EPA References:
U.S. EPA. 2018 Sampling, Laboratory and Data Consider-
1.1 This guide provides a framework for the application of
ations for Microbial Data Collected 32 the Field. EPA/
molecular biological tools (MBTs) to assess and characterize
600/R-164. 74 pg
in-situ biological processes to improve contaminated soil and
U.S. EPA. 2008 A Guide for Assessing Biodegradation and
groundwater management. While the focus of this guide is on
Source Identification of Organic Ground Water Contami-
in-situ biological processes, some concepts of how to apply
nants Using Compound Specific IsotopeAnalysis (CSIA).
MBTscanalsobeappliedtoex-situbioremediationapproaches
Ada, Oklahoma: Office of Research and Development,
(for example, biopiles, bioreactors) to support design,
U.S. (EPA 600/R-08/148)
operation, and troubleshooting. The intent of this guide is to
U.S. EPA. 1999 Use of monitored natural attenuation at
develop a consistent way in which MBTs are applied at
superfund, RCRA corrective action, and underground
contaminated sites, not to develop expertise. Technical experts
storage tank sites. United States Environmental Protection
need to be engaged whenscoping, planning, executing, and
Agency, Washington
interpreting data for MBTs. Lastly, there is a brief description
U.S. EPA 1996 Ground Water Issue: Low-Flow (Minimal
of isotopic techniques within section 5.2; however, the scope
Drawdown) Ground-Water Sampling Procedures EPA/
and focus of this guide is the use of nucleic acid-based MBTs
540/S-95/504
to assess biological processes at contaminated sites.
1.2 This standard does not purport to address all of the
3. Terminology
safety concerns, if any, associated with its use. It is the
3.1 This section includes definitions of key processes that
responsibility of the user of this standard to establish appro-
are specific to molecular biological tools and use of abbrevia-
priate safety, health, and environmental practices and deter-
tionsandacronyms.Definitionsareadaptedfromothersources
mine the applicability of regulatory limitations prior to use.
noted in the referenced documents. A full list of
1.3 This international standard was developed in accor-
bioremediation-relevant microorganisms is beyond the scope
dance with internationally recognized principles on standard-
of this guide.
ization established in the Decision on Principles for the
3.2 Definitions:
Development of International Standards, Guides and Recom-
3.2.1 16S ribosomal ribonucleic acid (rRNA), n—the RNA
mendations issued by the World Trade Organization Technical
component of the 30S unit of the prokaryotic ribosome. The
Barriers to Trade (TBT) Committee.
16S rRNA is produced by 16S rRNA genes (sometimes
referred to as 16S rDNA) which are gene sequences used to
2. Referenced Documents
study bacterial phylogeny and taxonomy.
2.1 ASTM Standards:
3.2.1.1 Discussion—The 16S rRNA gene has historically
D6771 Practice for Low-Flow Purging and Sampling Used
been the most common housekeeping genetic marker used by
for Groundwater Monitoring
molecular biologists and microbiologists to identify and clas-
sify microorganisms (for example, genus and species).
3.2.2 attenuation, n—contaminant reduction over space and
ThisguideisunderthejurisdictionofASTMCommitteeE50onEnvironmental
time due to physical (advection and dilution), chemical
Assessment, Risk Management and CorrectiveAction and is the direct responsibil-
(volatilization, adsorption, abiotic transformation), and/or bio-
ity of Subcommittee E50.04 on Corrective Action.
logical processes (biodegradation, biotransformation).
Current edition approved July 1, 2022. Published September 2022. DOI:
10.1520/E3354–22.
For referenced ASTM standards, visit the ASTM website, www.astm.org, or
contact ASTM Customer Service at service@astm.org. For Annual Book of ASTM AvailablefromUnitedStatesEnvironmentalProtectionAgency(EPA),William
Standards volume information, refer to the standard’s Document Summary page on Jefferson Clinton Bldg., 1200 Pennsylvania Ave., NW, Washington, DC 20460,
the ASTM website. http://www.epa.gov.
Copyright © ASTM International, 100 Barr Harbor Drive, PO Box C700, West Conshohocken, PA 19428-2959. United States
E3354 − 22
3.2.3 bioinformatics, n—a subdiscipline of biology and reaction typically of use or interest beyond its role in the
computer science that focuses on the acquisition, storage, general metabolism of the cell (housekeeping gene).
analysis and study of biological data, frequently applied to
3.2.15.1 Discussion—For example, the anaerobic Benzene
DNA, RNA, and protein sequences.
Carboxylase (abcA) gene encodes an enzyme responsible for a
known initial activation step in the anaerobic biodegradation
3.2.4 biomarker, n—a unique characteristic of a
pathway of benzene and is therefore a functional gene of
biomolecule, such as a specific DNA sequence, that can be
interest to bioremediation practitioners.
measuredandusedasanindicatorofatargetmicroorganismor
a specific biological process.
3.2.16 genomics,n—abranchofmolecularbiologyfocusing
3.2.5 bioaugmentation, n—the introduction of microorgan-
on the structure, function, evolution, and mapping of genomes.
isms into the environment for the purpose of enhancing a
3.2.16.1 Discussion—While genetics may focus on the
beneficial biological activity.
study of individual genes, genomics will consider the entire
3.2.6 bioremediation,n—cleanupofsitescontaminatedwith genetic information of an organism.
hazardous, toxic, or radioactive substances or wastes, or any
3.2.17 genome, n—the entire complement of genetic mate-
combination thereof using biological systems.
rial for an organism.
3.2.7 biostimulation, n—the introduction of electron donor,
3.2.18 metagenomics, or meta-omics, n—the study of ge-
electron acceptor, and/or nutrients to the subsurface to promote
netic material derived directly from multiple host organisms or
biodegradation by an existing, native microbial community.
environmental samples containing multiple genomes, such as a
Manipulation of pH or temperature may also be a form of
mixed microbial community.
biostimulation to enhance biodegradation.
3.2.19 molecular biological tools (MBTs), n—a suite of
3.2.8 compound specific isotope analysis (CSIA), n— ana-
molecular genetic analyses that can be used to characterize and
lytical method that measures the ratio between the stable
13 12 2 1 37 35
evaluate microorganisms and their related activity.
isotopes (for example, C/ C, H/ H, or Cl/ Cl) of a
3.2.19.1 Discussion—MBTs may also be referred to as or
contaminant. As contaminants are degraded (for example,
included in Environmental Molecular Diagnostics (EMDs).
metabolism of aromatics, reductive dechlorination of chlori-
nated solvents), contaminants become enriched with the
3.2.20 metabolism, n—the myriad of chemical reactions in
heavier isotopes (for example, C) and the associated shift in
organisms that support life, including the conversion of the
isotopic ratios can be determined as evidence of bond breaking
energy in substrates to energy available to perform cellular
reactions.
processes; the conversion of food to building blocks for
3.2.9 constituent of concern (COC), n—an environmental
proteins, lipids, nucleic acids, and some carbohydrates; and the
constituent that is to be assessed, remediated, and or monitored
elimination of metabolic wastes.
in groundwater, soil, or soil gas at contaminated sites; some-
3.2.21 metabolite, n—a low molecular weight intracellular
times referred to as a chemical of concern.
or extracellular molecule which is an intermediate or end
3.2.10 conceptual site model (CSM), n—a written and/or
product of biological processes required for organism growth,
graphical representation of a site which includes potential
maintenance, and normal function.
sources and receptors of contaminants as well as the collection
3.2.22 metabolic byproduct, n—biochemical compounds
of physical, chemical, and biological processes governing
that are generated by a microorganism during metabolic
contaminant fate and transport. These models are often
activity.
iterative, revised as more information is learned, and used to
aid decision-making throughout the site project lifecycle.
3.2.23 metabolomics, n—the analysis of intracellular and
extracellular intermediates and end products generated during
3.2.11 deoxyribonucleic acid (DNA), n—a biological mac-
biological processes required for growth, maintenance, and
romolecule that carries genetic information. DNA consists of
normal function of an organism.
two long chains of nucleotides twisted into a double helix.
3.2.12 electron acceptor, n—a chemical compound that
3.2.24 microcosm study, n—or “laboratory treatability stud-
accepts electrons transferred to it from another compound. It is
ies” are a laboratory study in which samples of environmental
an oxidizing agent that is reduced through the process of
media (for example, soil, sediment, and/or water) are tested in
accepting electrons.
a contained environment, such as a bottle, to evaluate changes
in concentration (or mass) of COCs over time.
3.2.13 electron donor, n—a chemical compound that do-
nateselectronstoanothercompound.Itisareducingagentthat 3.2.24.1 Discussion— Microcosm studies are designed to
is oxidized through the process of losing electrons.
mimicfieldconditions(totheextentpossible)andmaybeused
to evaluate remedial amendments, or determine natural degra-
3.2.14 enhanced attenuation (EA), n—a remedial strategy
dation rates.
that promotes biodegradation via addition of amendments
(biostimulation) or cultured microorganisms (bioaugmenta-
3.2.25 monitored natural attenuation (MNA), n—remedial
tion) that stimulate biogeochemical reactions that result in the
approach that relies on demonstration through multiple lines of
reduction of COCs in-situ.
evidence, that naturally occurring physical (advection and
3.2.15 functional gene, n—a segment of DNA that encodes dilution), chemical (volatilization, adsorption, abiotic
an enzyme or other protein that performs a known biochemical transformation), and/or biological processes (biodegradation,
E3354 − 22
biotransformation) are able to reduce the mass and concentra- 3.2.36 stable isotope probing (SIP), n—a technology that
tionofCOCstoachievesitespecificremedialobjectiveswithin utilizes a synthesized form of the contaminant containing a
stable isotope (for example, C label) added to a sampler or
a reasonable time frame.
study sample.
3.2.26 next generation sequencing (NGS), n—colloquial
3.2.36.1 Discussion—If biodegradation is occurring, the
term for massively parallel sequencing methods (for example,
isotope will be detected in the biomass (for example,
Illumina®) which allows the sequencing of millions of nucleic
phospholipids, DNA) and/or metabolic products (for example,
acid sequences from environmental samples simultaneously
volatile fatty acids, carbon dioxide or dissolved inorganic
and can be used for target amplicon sequencing or metag-
carbon or methane) over time.
enomic sequencing.
3.2.37 transcriptomics, n—analysis of all RNA transcripts
3.2.27 nucleobase (base), n—biological molecules that are
in a sample that are produced by the genome(s), under specific
cyclic organonitrogen compounds which are components of
circumstances, or in a specific cell using high-throughput
nucleotides that are part of fundamental structure of DNAand
methods, such as microarray analysis.
RNA.
3.2.38 transcripts, n—RNA synthesized from a DNA se-
3.2.27.1 Discussion—The five primary nucleobases are: ad-
quence (gene) that may then be translated in 295 proteins and
enine (A), cytosine (C), guanine (G), thymine (T), and uracil
ultimately the activation of a biochemical pathway (that is,
(U).
gene expression).
3.2.28 nucleotide, n—biological molecules that contain a
4. Significance and Use
nucleobase, a five-carbon sugar (for example, ribose (RNA) or
4.1 Contaminated sites subject to remediation are growing
deoxyribose (DNA)), and a phosphate group which make up
in complexity and associated remediation costs, presenting a
DNA and RNA.
challenge for managers of contaminated sites. The need to
3.2.29 primer, n—a short, chemically synthesized, single-
properly monitor, evaluate, and report remediation processes
stranded DNA sequence that is used to prime the polymerase
(including physical, chemical, and biological) characterizing
chain reaction to by complementary binding to a target gene
siteconditionsandcontaminantmassandattenuationiscritical
sequence a thereby initiating the amplification of a targeted for the evaluation and selection of effective remediation
segment of DNA. strategies. Assessment and characterization of biological pro-
cesses associated with contaminant attenuation is supported
3.2.30 prokaryote, n—unicellular organism that lacks a
and improved by the accurate and consistent use of molecular
membrane-bound nucleus. Prokaryotes are divided into two
biological tools (MBTs) including data acquisition,
domains, Archaea and Bacteria.
interpretation, and reporting.
3.2.31 proteomics, n—the analysis of the proteome, the
4.2 The development of this guide through ASTM Interna-
entire set or subset of proteins that is produced or modified by
tional is designed to meet the needs of managers of contami-
an organism or system during a biological process, such as
nated sites within the United States and elsewhere. The variety
contaminant transformation.
of available MBTs and the complexity with which they are
currently being applied are not addressed in existing ASTM
3.2.32 polymerase chain reaction (PCR), n—a laboratory
International Standards. The principal users of this guide
method used to make copies of a specific DNA sequence,
should be industry project managers, regulators, consultants,
allowingforamplificationtoasufficientscaleandresolutionto
analytical laboratories, and community stakeholders.
study and identify specific organisms and functional genes
3.2.33 quantitative polymerase chain reaction (qPCR), n—a 5. Overview of Molecular Biological Tools
laboratory analytical technique for quantification of a target
5.1 This guide provides an overview and suggested appli-
gene based on DNA polymerase chain reaction (PCR)
cation of a group of biological analyses collectively termed
technology, sometimes referred to as real-time PCR.
molecular biological tools (MBTs). MBTs are available to
environmental practitioners for the purpose of directly assess-
3.2.34 reverse transcriptase qPCR (RT-qPCR), n—a labo-
ing the contaminant-degrading capabilities and other activities
ratory analytical technique for quantification of an expressed
of microorganisms present in the environment. MBTs can be
target gene (RNA) based on production of complementary
applied to aqueous and solid media samples (for example,
DNA (cDNA) reverse transcribed from RNA.
groundwater, wastewater, surface water, soils, sediments, etc.).
3.2.35 ribonucleic acid (RNA), n—a single-stranded nucleic
There is a wide variety of MBTs, including but not limited to
acid that is transcribed from DNA and contains the comple-
quantitative polymerase chain reaction (qPCR) and
mentary genetic information.
metagenomics,thatcanbeapplieddependingontheselectivity
3.2.35.1 Discussion—Messenger RNA (mRNA) is trans-
desired (Fig. 1). For example, metagenomics allows for the
lated to amino acid sequences to synthesize proteins, including
sequencing on all DNA within a given sample, whereas target
enzymes responsible for biodegradation. Ribosomal RNA
amplicon sequencing only sequences genes selected for with
(rRNA) forms structural components of ribosomes which are primers often across all microorganisms containing a gene (for
required for protein synthesis. Transfer RNA (tRNA) is used
example, all bacterial 16S rRNA genes within a sample) and
during production of proteins in the ribosome. qPCR often selects a gene unique to microbial group or
E3354 − 22
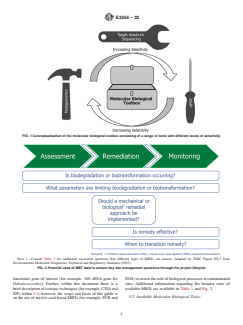
FIG. 1 Conceptualization of the molecular biological toolbox consisting of a range of tools with different levels of selectivity
NOTE 1—Consult Table 1 for additional associated questions that different types of MBTs can answer. Adapted by ITRC Figure ES-3 from
Environmental Molecular Diagnostics Technical and Regulatory Guidance (2013).
FIG. 2 Potential uses of MBT data to answer key site management questions through the project lifecycle
functional gene of interest (for example, 16S rRNA gene for NGS)toassesstheroleofbiologicalprocessesatcontaminated
Dehalococcoides). Further, within this document there is a sites. Additional information regarding the broader suite of
briefdescriptionofisotopictechniques(forexample,CSIAand
available MBTs are available in Table 1 and Fig. 3.
SIP) within 5.2; however, the scope and focus of this guide is
5.2 Available Molecular Biological Tools:
on the use of nucleic acid-based MBTs (for example, PCR and
E3354 − 22
TABLE 1 List of Molecular Biological Tools and Associated Questions, Type of Data Generated, Advantages and Disadvantages
Molecular Biological Type of Data
Abbreviation Associated Questions Advantages Disadvantages
Tool Generated
Polymerase Chain PCR Is a specific gene •Presence / absence •Culture independent •Positive results are dependent
Reaction (Targeted present? of gene •Sensitive upon sequence similarity to a
method) •Results are easy to understand known gene.
and convey to stakeholders •Does not give information on
•Inexpensive activity
Commercially available •Does not differentiate between
live and dead cell DNA
Quantitative qPCR How many copies of a •Presence / absence •Culture independent •Positive results are dependent
Polymerase Chain specific gene are of a gene •Sensitive upon sequence similarity to a
Reaction (Targeted present? •# of gene copies if •Quantitative known gene.
method) present •Results are easy to understand •Does not give information on
and convey to stakeholders activity
•Inexpensive •Does not differentiate between
•Commercially available live and dead cell DNA
Reverse Transcription RT-PCR Is a specific gene # of gene copies being •Culture independent •Positive results are dependent
Polymerase Reaction actively being transcribed •Quantitative upon sequence similarity to a
(Targeted method) transcribed (used) and Identifies genes that are actively known gene.
how much? transcribed •Activity is inferred from
•Results are easy to understand transcription
and convey to stakeholders •Only actively transcribed genes
•Inexpensive will be detected
•Commercially available
16S rRNAAmplicon 16S What microorganisms Relative percentages •Culture independent •Sensitivity is inversely
Sequencing sequencing are present? of distinct 16SrDNA •Provides information on what and proportional to complexity of the
(sometimes marketed (NGS) What microorganisms genes how many organisms are present community.
as Next Generation are numerically •Commercially available •Does not give information on
Sequencing) (Targeted dominant? activity
method) •Does not differentiate between
live and dead cell DNA
•Bioinformatic analysis can be a
bottle neck to data usability and
conveying results to stakeholders
Metagenomics (Non- What organisms and Assemblage of the •Culture independent •Does not give information on
targeted method) functional genes are genetic material of the •Commercially available activity
present in the sampled entire microbial •Does not differentiate between
microbial community? community at a live and dead cell DNA
specific point in time •Bioinformatic analysis can be a
bottle neck to data usability and
conveying results to stakeholders
Transcriptomics (Non- •What genes are being Assemblage of the •Culture independent •Bioinformatic analysis can be a
targeted method) used by the microbial genes being actively •Provides information on which bottle neck to data usability and
community? transcribed of the physiological processes should be conveying results to stakeholders
•Do they change entire microbial active •Not commercially available
under different community at a
conditions? specific point in time
Proteomics (Non- What proteins/ Assemblage of the Culture independent Requires specialized expertise for
targeted method) enzymes are being proteins being Can be used to reverse engineer separation and interpretation
produced by the produced by of the a nucleic acid probe for a gene of Not commercially available
microbial community? entire microbial interest
community at a Provides information on what
specific point in time processes organisms are actively
carrying out in the environment.
Stable Isotope Probing SIP Is a specific target a) Shift of DNA to •Can absolutely confirm •Limited to compounds that are
(addition of stable compound being higher density biodegradation of target used as sources of carbon by
isotope) biodegraded under in- indicating uptake compound under in-situ conditions microbial community
situ conditions? b) Identification of •When used with passive
degrading community samplers the size of the microbial
through higher density population may differ from the
DNA in 16SrDNA actual environment due to
sequence fragments selective enrichment in the
sampler.
Compound Specific CSIA Is compound Change in ratio of •Can confirm degradation of target •Limited to degradation reactions
Isotope Analysis attenuation due to heavy to light compound under in-situ conditions that result in a strong isotope
(natural abundance of destructive reactions? isotope(s) in the (as opposed to non- degradative fractionation.
stable isotope (no compound of interest attenuation mechanisms). •Detection limits for stable isotope
addition) relative to absolute •Can provide information about are typically much higher than for
concentration of the the primary mechanism (biotic/ mass measurements.
compound. abiotic) of target compound •Significant method validation
destruction in a sample. required for each compound of
interest.
Metabolomics What processes are Assemblage of the Can provide information on the •Requires specialized expertise
active in the microbial metabolite profile of a active physiological processes in for separation and interpretation
community? cell or community at a the environment •Not commercially available
specific point in time
E3354 − 22
NOTE 1—Adapted from schematic on “Metabolome” Wikipedia page (https://en.wikipedia.org/wiki/Metabolome).
FIG. 3 Conceptual schematic depicting overview and relationship between select MBTs, biological ensemble, biological molecules
(DNA, RNA, proteins, and metabolites), and type data generated
5.2.1 Polymerase Chain Reaction: tion that may result from environmental contamination and
5.2.1.1 The primary MBTs described in this guide are those remediation activities. NGS is not a fully quantitative tool, but
based on the polymerase chain reaction (PCR). The goal of rather provides insight on the proportions or abundances of
PCR analysis is the amplification of a target nucleic acid microbes relative to others in a given community. Recent
sequence (genomic DNA [gDNA] or complementary DNA advances in sequencing technology have significantly reduced
[cDNA])byseveralordersofmagnitude.Onceamplified,there the cost of this analysis. Nevertheless, the organization and
should be sufficient copies of DNA for further analysis and analysis of the sequenced DNA requires bioinformatic exper-
interpretation. Through the amplification process, PCR tech- tisewhichcanbeabottlenecktoproducingandinterpretingthe
nologies allow detection and analysis of DNAsequences even results. NGS technology can be used for targeted amplicon
whentheyareinitiallypresentatlowlevelsintheenvironment. sequencing,suchasforthe16SrRNAgene,orfornon-targeted
5.2.1.2 Many adaptations have been made to this technique, sequencing of the entire metagenome (metagenomics) within a
including reverse transcription PCR (RT-PCR), quantitative sample. When requesting NGS from a laboratory, it is impor-
real-time PCR (qPCR), reverse transcription qPCR (RT-qPCR) tant to specify targeted amplicon or metagenomic sequencing
and digital PCR (dPCR). In all cases, the PCR analysis relies and understand laboratory NGS 382 capabilities.
on the use of a synthesized target sequence of DNA (that is, 5.2.3 Omics—Omics is a term that broadly refers to the
primers) to query total genomic DNA extracted from environ- study of biological processes at numerous levels, from the
mental samples for the presence of complementary sequences. characterization of genes (genomics), mRNA
5.2.1.3 qPCR, allows for a targeted gene to not only be (transcriptomics), proteins (proteomics) or metabolites (me-
detected but also quantified.This technology is regarded as the tabolomics) (see Fig. 3). While genomics is the study of an
“gold standard” nucleic acid analysis for contaminated site organism’s entire genetic material, metagenomics is the study
samples because of its high sensitivity, good reproducibility, of genetic material of all of the organisms found in a specific
quantification range, ease of use and relatively low costs. environment. In the context of contaminated site management,
Quantification of a target DNA sequence (over time or space) these tools can provide yet another line of evidence to support
using qPCR allows for evaluation of trends related to key the activity of specific physiological functions (for example,
organismsorfunctionalgenesinvolvedincontaminantbiodeg- biodegradation of hydrocarbons or sulfate reduction), or docu-
radation. ment changes or differences in microbial populations.
5.2.2 Next Generation Sequencing—Next Generation Se- 5.2.4 Isotopic Tools—Isotopic tools rely on measurable
quencing (NGS) is the simultaneous sequencing of multiple differencesbetweenheavyandlightstableisotopesofelements
pieces of nucleic acid for the purpose of making observations in a single compound.
about the full microbial community within a sample. This 5.2.4.1 Compound specific isotope analysis—Compound
technology allows for assessment of the microbial community specific isotope analysis (CSIA) measures differences in rela-
(that is, phylogenetics or who is there) and, in some cases tive abundance of stable isotopes within a COC, for example
13 12
wherefunctionisstronglylinkedtophylogeny,suggestorinfer C and C, especially when chemical bonds are expected to
what the microbes do (for example, their function or role). be broken. For remediation strategies where COC mass is
NGS can be a useful tool to observe changes in microbial transformed or destroyed (through chemical transformation or
community structure both in microbial diversity and in func- biodegradation), the bonds in lighter isotopes (for example,
E3354 − 22
C) are preferentially broken resulting in an enrichment of the amendments (for example, emulsified vegetable oil (EVO),
heavier isotopes (for example, C) of the remaining parent formate, sulfate, nitrate, oxygen, etc.). Alternatively, the key
substrate. These compound specific isotope changes can be
microorganisms may not be present in sufficient numbers
linked to the extent of transformation or biodegradation of because the site geochemistry is not supportive. For this latter
COC over time or distance (USEPA, 2008).
case, the remedial strategy could be changing the overall
5.2.4.2 Stable isotope probing—Stable isotope probing
geochemistry, for example pH, and possibly bioaugmentation,
(SIP) uses a microbial substrate (typically a site COC which is
the addition of key microorganisms.
enriched in the heavier stable isotope and can be purchased
6.1.3 Monitoring—MBTs can be a powerful tool for moni-
from a specialized vendor) The heavy isotope contaminant is
toring the progression of an implemented remedy. MBTs can
subjected to the site subsurface conditions and microbes that
directly measure the presence of key microorganisms, func-
then may degrade the heavier contaminant. Biomass and
tional genes that code for enzymes involved in biodegradation
biodegradation products are tracked over time to assess incor-
of COCs, and other relevant biogeochemical processes. Moni-
poration of the heavier isotope to demonstrate ongoing biodeg-
toring of microbial and functional genes, in conjunction with
radation processes. SIP can be performed in the lab and in the
changes in COC concentrations and geochemical data, often
field using commercially available in-situ microcosm devices.
provide compelling evidence to determine the effectiveness of
remedial interventions. For example, MBT data may demon-
6. Application of Molecular Biological Tools
strate an increased abundance of contaminant-degrading mi-
6.1 Project Lifecycle—The application of MBTs should be
croorganismsorfunctionalgenesbeforemeasurablechangesin
considered for use during the following stages of the contami-
contaminant concentration are observed. These data are espe-
nated site project lifecycle: assessment, remediation, and
cially relevant when there is significant contaminant mass
monitoring. Application of MBTs can be performed to answer
present as sorbed phase or non-aqueous phase liquid (NAPL)
different questions depending on the stage of the project
in the area targeted by remediation. This observation may be
lifecycle in which MBTs are applied (Fig. 2). MBTs can be
considered a leading indicator of contaminant remediation. If
complementary lines of evidence used to: (a) develop or refine
required microorganisms are not detected at sufficient
biogeochemical processes within the conceptual site model
abundance, that finding may indicate the need for remedy
(CSM) (b) reduce uncertainty of biological processes associ-
adjustment or optimization.
ated with remedy design, (c) monitor remedy performance and
6.2 Value of Application:
differentiate biological processes from chemical or physical
processes, and (d) support communications with stakeholders.
6.2.1 Assessing Contaminant Biodegradation—The com-
While nucleic acid-based MBTs cannot quantify field-scale
mercial availability of PCR and qPCR technologies has re-
biodegradation or biotransformation rates as of the time of
sulted in their routine use for detecting presence or absence of
writing this guide, there are past and ongoing efforts to assess
specific microorganisms or functional genes known to biode-
the potential for field-scale rate quantification based on the
grade COCs and in the case of qPCR, quantification. For
abundance of biomarkers (Lu et al., 2006 (1), Wilson et al.,
example, high abundance of Dehalococcoides mccartyi (an
2019 (2), Michalsen et al., 2021 (3),Adamson et al., 2022 (4).
obligate organohalide respiring anaerobe) and vinyl chloride
6.1.1 Assessment—MBT data in conjunction with COC, or
reductase genes serve as positive indicators of the potential for
geochemical data, can be used to support site assessment by
anaerobic biodegradation of chlorinated organics. As addi-
determining if key organisms or genes associated with biodeg-
tional COC biodegradation pathways, and corresponding asso-
radationarepresentatthesite.WiththeMBTandgeochemical
ciated functional genes, are identified, qPCR assays are devel-
data set, a project manager can determine if the baseline site
oped to target these novel genes. Currently, there are well
conditions are conducive to biodegradation processes. If COC
documented qPCR assays to quantify microorganisms and
attenuationhasbeenobserved,thesedatacanhelpestablishthe
functional genes that degrade a wide range of environmental
mechanisms responsible for observed changes. If COC attenu-
contaminants. Table 2 lists a selection of some of the commer-
ation has not been observed, these data can help in the
cially available qPCR test targets as of October 2021. It is
development of hypotheses as to the factors limiting biodeg-
inevitable that as additional contaminant biodegradation and
radation.
biotransformation pathways, microorganisms, and functional
6.1.2 Remediation—Once the site has been assessed at the
genesareidentifiedandelucidated,newassayswillcontinueto
baseline phase, MBT data can support the selection and
be developed and deployed.
implementation of specific remedial strategies to initiate or
6.2.2 Supporting Remedy Transition—Various remediation
enhance attenuation of COCs. For example, COC attenuation
technologies are often combined to remove contaminant mass
could be occurring slowly or have stalled due to a depletion of
through physical processes (for example, volatilization or
limitingelectrondonors,electronacceptors,ornutrientsevenif
dissolution from NAPL) and then destroy it via chemical
key microorganisms are present. In this case, the remedial
and/or biological processes (for example, chemical oxidation
strategy might be to alleviate the electron acceptor/donor or
and biodegradation). Mechanical in-situ remediation ap-
nutrient limitation through biostimulation with site-specific
proaches include technologies such as pump-and-treat, air
sparging, soil vapor extraction, or multiphase extraction for
4 contaminant removal from saturated and/or vadose zone. At
The boldface numbers in parentheses refer to a list of references at the end of
this standard. sites being treated with mechanical processes, the remediation
E3354 − 22
TABLE 2 Selection of Commercially Available Genetic Targets for Microorganisms and Functional Genes Associated with Contaminant
Biological Processes
Contaminant Class Redox Acronym Target Most Common Relevance
Chlorinated Ethenes Anaerobic Dhc Dehalococcoides Reductively dechlorinates PCE, TCE, all DCE isomers,
VC
bvcA BAV1 Vinyl chloride reductase (bvcA) Dechlorination of cDCE and VC to ethene
tceA Trichloroethene reductase (tceA) Dechlorination of PCE and TCE to cDCE and VC
vcrA Vinyl chloride reductase (vcrA) Dechlorination of cDCE and VC to ethene
Dhb Dehalobacter Partial dechlorination of PCE and TCE to cDCE
Dsm Desulfuromonas Reductively dechlorinates PCE and TCE to cis-DCE
using acetate as an electron donor
Dsb Desulfitobacterium Reductively dechlorinates PCE and TCE to cis-DCE
PCE-1 Geobacter Tetrachloroethylene Reductive dechlorination of PCE to cis-DCE
Reductase (pceA)
PCE-2 Sulfurospirillum Tetrachloroethylene Reductive dechlorination of PCE to cis-DCE
Reductase (pceA)
Dhg Dehalogenimonas Dechlorination of tDCE to VC and VC to ethene
CER Dehalogenimonas Vinyl Chloride Dechlorination of VC to ethene
Reductase
TDR Dehalogenimonas 1, 2-Trans Reductive dechlorination of 1,2 trans DCE
Dichloroethene Reductive
Dehalogenase (tdrA)
MBR DHC MB Reductase Potential biomarker for Production of trans DCE during
dechlorination of PCE and TCE
MGN Methanogens via methyl coenzyme Utilize hydrogen and can compete with halorespiring
reductase mcrA/mrtA bacteria for available electron donor
APS Sulfate reducing bacteria via Compete with halorespiring bacteria for available
Adenosine 5’ phosphosulfate hydrogen
reductase
Aerobic sMMO Soluble Methane Monooxygenase Co-oxidation of TCE, cis-DCE, and vinyl chloride.
Expressed at higher copper to biomass ratios
pMMO Particulate Methane Monooxygenase pmoA is expressed at higher copper to biomass ratios
and is capable of cometabolizing TCE at lower rates.
RMO Ring hydroxylating Toluene Co-oxidation of TCE
monooxygenase (toluene-3 and 4
monooxygenases)
RDEG Ring hydroxylating Toluene Co-oxidation of TCE
monooxygenase (toluene-2
monooxygenases)
PHE Phenol Hydroxylase Co-oxidation of TCE
TOD Toluene and Benzene Dioxygenases Co-oxidation of TCE
EtnC Ethene monooxygenase Converts ethene and vinyl chloride to their respective
epoxyalkanes
EtnE Epoxyalkane transferase Mediates conjugation and breaking of the epoxide
PPO Propane monooxygenase Cometabolism of TCE in the presence of propane
BMO Butane monooxygenase Cometabolism of TCE in the presence of butane
Chlorinated Ethanes Anaerobic Dhb Dehalobacter Dechlorination of 1,1,1-TCA/1,1-DCA to chloroethane;
1,2-DCA /1,1,2-TCA to VC/ethene; 1,1,2,2-TeCA to tDCE
Dhg Dehalogenimonas Dechlorination of 1,2- DCA to ethene; 1,1,2,2-TeCA to
c/tDCE; 1,1,2-TCA to VC
Dhc Dehalococcoides Dechlorination of 1,2-DCA to ethene; dechlorination of
VC produced from 1,1,2-TCA dechlorination
Dsb Desulfitobacterium Dechlorination of 1,1,2-TCA and 1,2-DCA
DCA 1,1 Dichloroethane dehalogenase Dechlorination of 1,1-DCA to chloroethane
(dcrA)
CFR Chloroform Reductase (cfrA, ctrA, dechlorination of 1,1,1-TCA to 1,1-DCA
thmA)
DCAR 1, 2 Dichloroethane dehalogenase Dechlorination of 1,2-DCA to ethene
(Desulfitobacterium and Dehalobacter)
MGN Methanogens via methyl coenzyme Utilize hydrogen and can compete with halorespiring
reductase mcrA/mrtA bacteria for available electron donor
APS Sulfate reducing bacteria via Compete with halorespiring bacteria for available
Adenosine 5’ phosphosulfate hydrogen
reductase
Aerobic sMMO Soluble Methane Monooxygenase Co-oxidation of 1,1,1-TCA and 1,2-DCA
PPO Propane monooxygenase Cometabolism of 1,1,2-TCA in the presence of propane
BTM Butane monooxygenase Cometabolism of 1,1,1-TCA in the presence of butane
Chlorinated Methanes Anaerobic Dhb Dehalobacter Reductive dechlorination of chloroform to DCM;
fermentation of DCM to acetate
DCM Dehalobacter DCM Degrade DCM through fermentation
CFR Chloroform Reductase (cfrA) CFR converts chloroform to dichloromethane
Aerobic sMMO Soluble Methane Monooxygenase Co-metabolism of chloroform and dichloromethane
DCMA Dichloromethane dehalogenase Dechlorination of DCM to formaldehyde under oxic
conditions
Chlorinated Propanes Anaerobic Dhg Dehalogenimonas Converts TCP to allyl chloride; DCP to propene
Dhc Dehalococcoides Converts DCP to propene
Dhb Dehalobacter Converts DCP to propene
E3354 − 22
TABLE 2 Continued
Contaminant Class Redox Acronym Target Most Common Relevance
Dsb Desulfitobacterium Dechlorination of TCP and DCP
1,2 DCP 1,2 Dichlorpropane Dehalogenase Dechlorination of 1,2 DCP to
Aerobic sMMO Soluble Methane Monooxygenase Capable of co-oxidation of 1,2-DCP 1,3-DCP and 1,2,3-
TCP
PPO Propane monooxygenase Capable of cometabolism of TCP
Chlorinated Benzenes Anaerobic Dhc Dehalococcoides Capable of Dechlorinating hexachlorobenzene,
pentachlorobenzene, tetrachlorobenzene
Dhb Dehalobacter Potential reductive dechlorination of DCBs and CB
DECO Dehalobium chlorocoercia DF-1 Capable of reductive dechlorination of HCB, PeCB and
1,2,3,5- TeCB.
MGN Methanogens via methyl coenzyme Utilize hydrogen and can compete with halorespiring
reductase mcrA/mrtA bacteria for available electron donor
APS Sulfate reducing bacteria via Compete with halorespiring bacteria for available
Adenosine 5’ phosphosulfate hydrogen
reductase
Aerobic PHE Phenol Hydroxylase Significant increases in numbers of bacteria containing
PHE genes corresponded to increases in biodegradation
of DCB isomers
TOD Toluene and Benzene Dioxygenases Mediates the incorporation of both atoms of oxygen into
the aromatic ring of benzene and substituted benzenes
(toluene and chlorobenzene)
RMO Ring hydroxylating Toluene Catalyze the initial and in some cases second oxidation
monooxygenase (toluene-3 and 4 of a variety of monoaromatic compounds including BTEX
monooxygenases) and CB
TCBO Trichlorobenzene and Biphenyl/ Capable of aerobic biodegradation of a number of
Isopropylbenzene Dioxygenases chlorinated benzenes including chlorobenzene, 1,2-
dichlorobenzene, 1,2,4- trichlorobenzene, and 1,2,4,5-
tetrachlorobenzene
Chlorinated Biphenyls Anaerobic Dhc Dehalococcoides Capable of reductive dechlorination of PCBs
DECO Dehalobium chlorocoercia DF-1 Shown to reductively dechlorinate doubly flanked
chlorines in PCBs
PCBR Dehalococcoides PCB Reductase May serve as biomarkers for the dechlorination of PCBs
pcbA1, pcbA4, and pcbA5
Aerobic BPH Biphenyl Dioxygenase Initiates aerobic cometabolism of PCBs.
Chlorinated Phenols Anaerobic Dhc Dehalococcoides Capable of utilizing PCP and tetrachlorophenol (TeCP),
trichlorophenol (TCP), and 2,3-dichlorophenol (2,3-DCP)
Dsb Desulfitobacterium Some strains are capable of reductive dechlorination of
PCP while others can utilize TCP and DCP
MGN Methanogens via methyl coenzyme Utilize hydrogen and can compete with halorespiring
reductase mcrA/mrtA bacteria for available electron donor
APS Sulfate reducing bacteria via Compete with halorespiring bacteria for available
Adenosine 5’ phosphosulfate hydrogen
reductase
Aerobic PCP Pentachlorophenol Monooxygenase Initiates oxidation of PCP and aromatic ring cleavage
(pcpB, E, and R)
BTEX Anaerobic bssA Benzylsuccinate Synthase Mediates the first step in the anaerobic biodegradation of
toluene and in some cases ethylbenzene and xylenes
abcA Anaerobic Benzene Carboxylase Anaerobic biodegradation of benzene
GMET Geobacter metallireducens Functional Targets functional genes including a predicted
Genes oxidoreductase specifically required for anaerobic
benzene metabolism
BCR Benzoyl Coenzyme A Reductase Reduces the benzene ring structure of the central
intermediate Benzyl-CoA
ORM-2 Deltaproteobacterium ORM-2 ORM-2 anaerobic benzene degrader under sulfate
reducing or methanogenic conditions
SRB Sulfate reducing bacteria via SRB are symbiotic partners to ORM-2 in anaerobic
dissimilatory sulfate reductase (dsrA) benzene degradation
Pepto-ben Benzene degrading Peptococcacae Anaerobic benzene degradation under nitrate reducing
conditions
Aerobic TOD Toluene and Benzene Dioxygenases Capable of degrading toluene, benzene, and
chlorobenzene along with co-oxidation of a variety of
compounds including ethylbenzene, p-xylene, and
m-xylene
RMO Ring hydroxylating Toluene Catalyze the initial oxidation and sometimes second
monooxygenase (toluene-3 and 4 oxidation steps in aerobic BTEX biodegradation
monooxygenases)
RDEG Ring hydroxylating Toluene Catalyze the initial oxidation and sometimes second
monooxygenase (toluene-2 oxidation steps in aerobic BTEX biodegradation
monooxygenases)
PHE Phenol Hydroxylase Catalyze the continued oxidation of phenols produced by
RMOs
TOL Toluene /Xylene Monooxygenase Biodegradation of toluene by attacking at the methyl
group
EDO Ethylbenzene Dioxygenase Responsible for aerobic biodegradation of alkylbenzenes
including ethylbenzene and isopropylbenzene or cumene
E3354 − 22
TABLE 2 Continued
Contaminant Class Redox Acronym Target Most Common Relevance
TCBO Trichlorobenzene and Biphenyl/ Includes benzene and isopropylbenzene dioxygenases
Isopropylbenzene dioxygenases
MTBE and TBA Aerobic PM1 Methylibium petroleiphilum PM1 Capable of utilizing MTBE and TBA as growth supporting
substrates
TBA Tert Butyl Alcohol Monooxygenase Catalyzes the continued biodegradation of TBA, an
intermediate produced during aerobic MTBE and ETBE
biodegradation
ETHB P450 cytochrome monooxygenase Initiates aerobic biodegradation of ETBE and is capable
of co- oxidation of MTBE and TAME
Diesel, Naphthalene, Anaerobic ANC Anaerobic Naphthalene Carboxylase Only known pathway for anaerobic biodegradation of
and PAHs naphthalene
mnssA Naphthyl-2-methyl-succinate Synthase Initiates anaerobic biodegradation of
2-methylnaphthalene by catalyzing the addition of
fumarate onto the methyl group
Aerobic NAH Naphthalene Dioxygenase Initiates aerobic metabolism of naphthalene by
incorporating both atoms of molecular oxygen into the
ring. Also capable of catalyzing oxidation of anthracene,
phenanthrene, acenaphthylene, acenaphthene, and
fluorine.
NidA Naphthalene-Inducible Dioxygenase Capable of mineralizing naphthalene and degrading
some higher molecular weight PAHs including pyrene
and benzo[a]pyrene.
PHN Phenanthrene Dioxygenase Phenanthrene/naphthalene dioxygenases capable of
degrading phenanthrene and naphthalene but have
broad specificity
ARHA Acenaphthylene Dioxygenase Capable of catalyzing the degradation of acenaphthene,
acenaphthylene, naphthalene, phenanthrene, anthracene
and fluoranthe
...



Questions, Comments and Discussion
Ask us and Technical Secretary will try to provide an answer. You can facilitate discussion about the standard in here.
Loading comments...